At Spring, we've dedicated ourselves to building a technology engine that helps those searching for therapies for many different diseases. This engine is capable of generating strong conviction in screening hits using complex, data-rich assays across thousands of drugs in a broad range of therapeutic areas. (Reach out to our partnerships team to learn more.)
One such therapeutic area that we've extensively validated is immune aging-related diseases.
Age-related diseases cause a tremendous amount of suffering. They are many, they are awful, and they don’t discriminate. Scientists work tirelessly to find therapies, but they need new tools – tools that give them the superpowers such a fight demands.
We've used our engine to discover therapies for certain diseases of immune aging. Our engine maps changes in our immune system, searches thousands and thousands of potential treatments, and gives scientists the tools to understand them.
Building and using such a tool for finding novel drugs requires:
- New technology to profile and perturb a wide array of immune biology.
- A non-traditional approach to drug discovery, placing engineered biological tools in the hands of scientists to help steer towards the most promising hypotheses.
- Validation across many diseases to prove the engine’s broad therapeutic relevance.
Running a multi-disease therapeutic engine
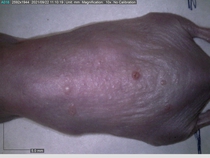
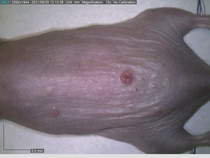
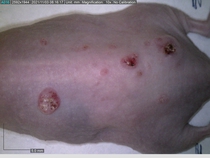
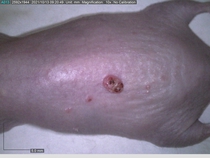
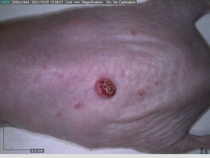
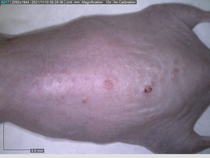
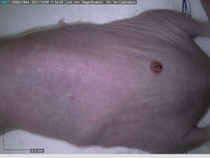
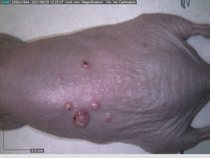
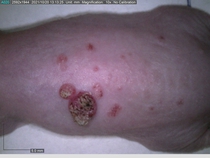
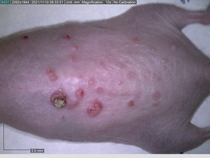
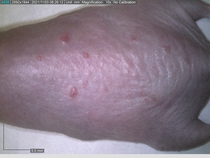
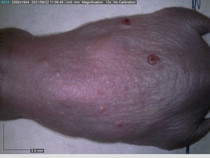
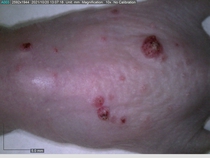
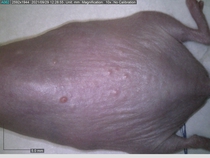
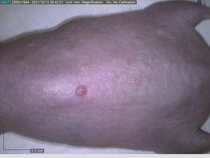
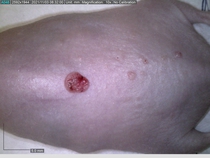
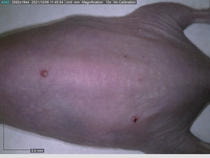
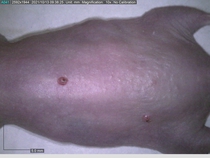
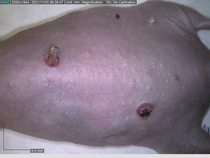
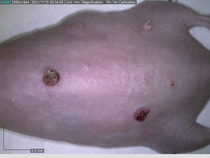
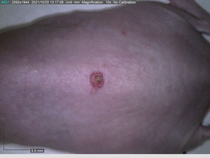
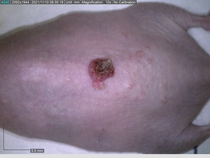
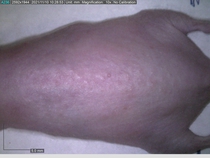
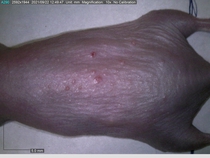
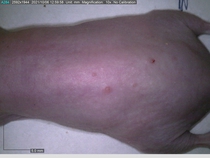
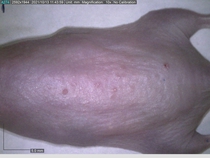
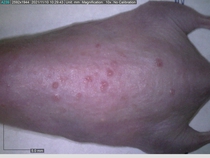
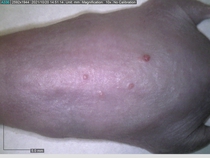
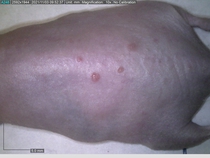
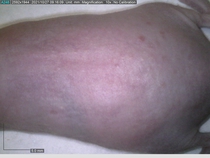
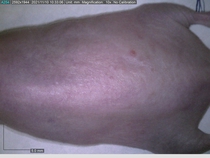
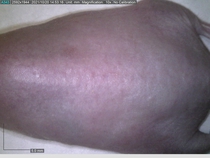
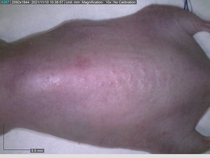
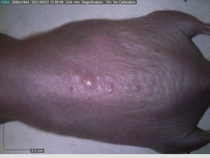
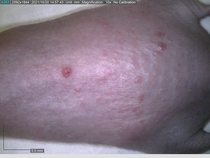
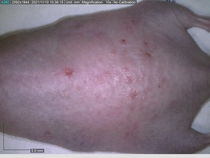
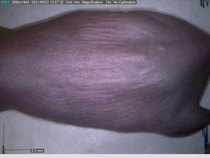
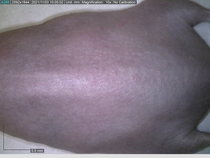
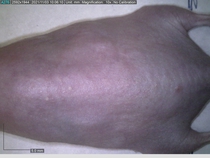
Starting with one example of the engine’s output: drugs to reduce the precancerous skin lesions that form after a lifetime of sun exposure.
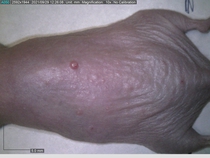
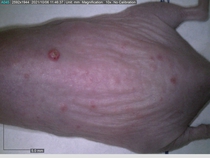
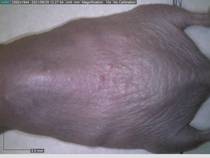
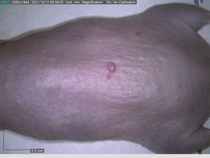
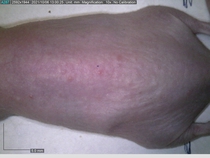
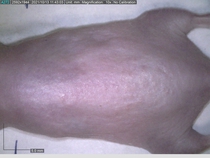
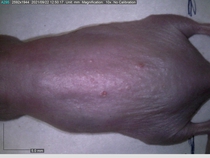
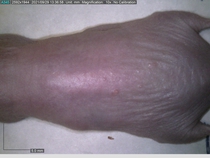
In a recent Spring study, multiple drugs evaluated by our novel screening technology and surfaced via MegaMap helped cure mice of these sorts of precancerous skin lesions associated with sun exposure. The treatment group experienced drastically fewer and smaller lesions, along with improved skin health overall.
This study is an example of the kind of follow-up validation often run at Spring – orthogonal, established protocols used to test the outputs of our engine. We run such validation experiments for our partners for many different diseases (more below).
But even more importantly, the hypotheses tested in this study weren't generated through a traditional approach to drug discovery.
Instead, they came from putting engineered biological tools in the hands of scientists – tools designed to help them wield enormous biological data, answer hard questions quickly, and surface the most promising hypotheses. Equipping our scientific partners with such superpowers creates repeatable conditions for promising drug discovery – an engine capable of addressing multiple diseases.
An engine should run again and again and again
Typical experiments in drug discovery measure very narrow ways in which drugs may or may not be helpful for affecting a specific disease target. This reductivist approach works really well, when it works – when we know the right targets and have strong conviction in their clinical relevance. But these aren’t easy conditions to recreate again and again – and certainly not for complicated diseases.
Aging and immune function, for example, are two incredibly complex, multi-faceted biological processes. Any singular measurement focused on an individual target or disease mechanism will fail to capture anything close to a complete picture of the biology.
This makes it especially challenging to apply the reductivist approach – and presents an opportunity to use new tools to search for clinically relevant therapies. Building these new tools for the field of immune aging has opened up a powerful therapeutic search process.
A landscape of novel immune therapies, using known drugs as waypoints
Using Spring's engine, we started by profiling a multitude of high-level immune cell behaviors that point towards therapeutic applications for multiple disease pathways – creating a rich, high-dimensional landscape of relevant biology for scientists to explore.
We then profiled thousands of therapies within that landscape of relevant biology – placing multiple waypoints along the way using known compounds and cell perturbations.
controls
in follow-up assay.
activation
controls
in animal study.
in follow-up assay.
Such reference profiles help guide our search towards areas of therapeutic interest – for example, therapies that activate the immune system, curb inflammation, or bolster vaccine response. By searching for unknown drugs that ‘look like’ these known experimental waypoints, we discover both novel drugs and novel mechanisms.
Combined with the phenotypic detail surfaced by MegaMap, our approach gives our research partners a compass to navigate potential therapies’ impact on complex immune function – uncovering drugs that induce the immune responses most relevant to their clinical interest, whether it be precancerous lesion formation, tumor growth, vaccine response, or another indication.
Validating efficacy in follow-up experiments for multiple diseases
The repeatability of this engine has allowed scientists to find promising drugs for multiple diseases. But validating their efficacy requires gold-standard experimental data – testing the outputs of our engine using independent, established protocols.
The design of these experiments varies from disease to disease: one, as seen above, involved exposing mice to UV to accelerate skin aging; another, seen below, involved using model vaccines to assess effects on vaccine efficacy.
In each disease case, we validated efficacy for our top drugs -- each of which was surfaced by the same, repeatable immune aging engine.
From decreasing tumor size to reducing skin lesions to improving vaccine response to maturing immune cells to decreasing proinflammatory cell death, this suite of clinically-relevant validation results have strengthened our belief in the multi-disease approach of our engine – and its applicability to many different therapeutic areas.
More fuel in the engine
We're now working with more partners, ramping up the number of therapeutic programs running on this engine, in collaboration with researchers around the world – while making non-stop investments in the software and technology behind it.
We strongly believe that scientists who are searching for treatments for disease are one of our greatest forces for good. And that equipping them with world-leading tools is one of the best possible uses of technology.
If you're one of these folks and want to be armed with the technology you deserve, contact us. We'd be excited to help.
Thanks to Jeffrey Chan for the disease model illustrations.